Dear NetLogo Community and Fellow Researchers,
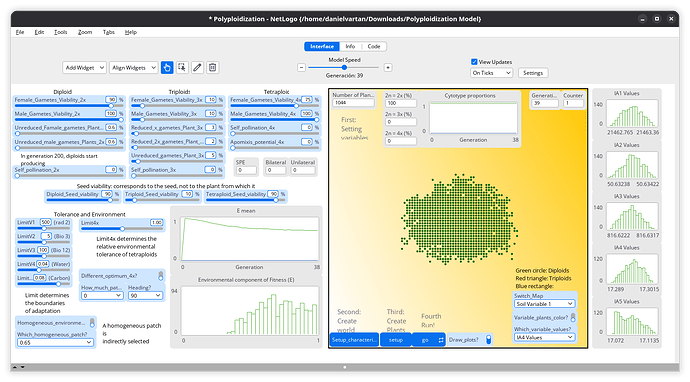
I am delighted to share a new agent model, designed to investigate the complex dynamics of diploid-polyploid interactions and the establishment of neopolyploids.
Model: Polyploidization model - Google Drive
Repository: http://hdl.handle.net/11336/253196
Article: https://doi.org/10.1038/s41598-025-29286-7
This model serves as a scalable, easy-to-use command center for analyzing the spatio-temporal dynamics of these evolutionary systems. It incorporates a multilayered environment where eco-variables fluctuate across generations, making the simulation highly responsive to environmental heterogeneity.
Key mechanisms within the model include the use of reproductive syndromes (like self-fertility and apomixis) and traits such as adaptivity, niche breadth, and dispersal.
Our preliminary analyses using this framework have provided clear, model-based evidence on several critical aspects: a large proportion of polyploidization events are unsuccessful, but factors like self-fertility, apomixis, and parental traits significantly accelerate establishment and demographic success. Furthermore, ecological niche shifts are shown to be vital for promoting cytotype coexistence.
My hope is that this NetLogo model will prove to be a valuable resource for your academic work, research projects, and teaching in the field of evolutionary and ecological modeling.
Please feel free to test the model, utilize it in your own research, and share any feedback or potential extensions you might consider. Your collaboration is highly valued.
Best wishes,
Juan Sebastián Schneider